Google Release: DeepVariant AI For Genomics
by TeachThought Staff
Google has announced the release of a new technology to map human genome sequences.
By outward appearances, this is a very narrow and niche application of technology that doesn’t have anything to do with 21st-century teaching and learning, a lesson here is more relevant for education: technology is most effective when designed expressly for a task, but it’s also true that one technology can be ‘pivoted’ for previously unconsidered uses.
Students using an iPad to operate apps? Sure.
Students using an iPad to document home life or create ‘closed-circuit’ documentaries? Doesn’t seem like the best ‘fit’ for a tablet, but tablets are mobile and students are mobile, so what is possible if we design learning as mobile-first through mobile-first teaching.
See also 10 Uses For Artificial Intelligence In Education
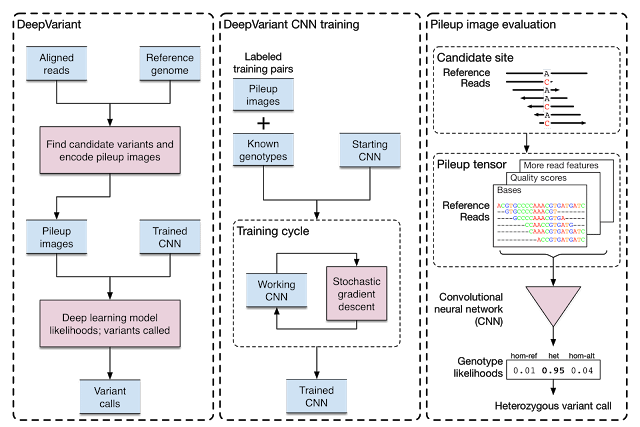
“Today, we announce the open source release of DeepVariant, a deep learning technology to reconstruct the true genome sequence from HTS sequencer data with significantly greater accuracy than previous classical methods. This work is the product of more than two years of research by the Google Brain team, in collaboration with Verily Life Sciences. DeepVariant transforms the task of variant calling, as this reconstruction problem is known in genomics, into an image classification problem well-suited to Google’s existing technology and expertise.”

The announcement continues, “DeepVariant is being released as open source software to encourage collaboration and to accelerate the use of this technology to solve real world problems….This is all part of a broader goal to apply Google technologies to healthcare and other scientific applications, and to make the results of these efforts broadly accessible.’
In this case, the trend—use new technology to tackle problems that previously were previously under-attended to because the technology simply didn’t exist—is interesting, and a useful lesson for education.
You can read more on Google Research Blog and on Wired.com for more background.